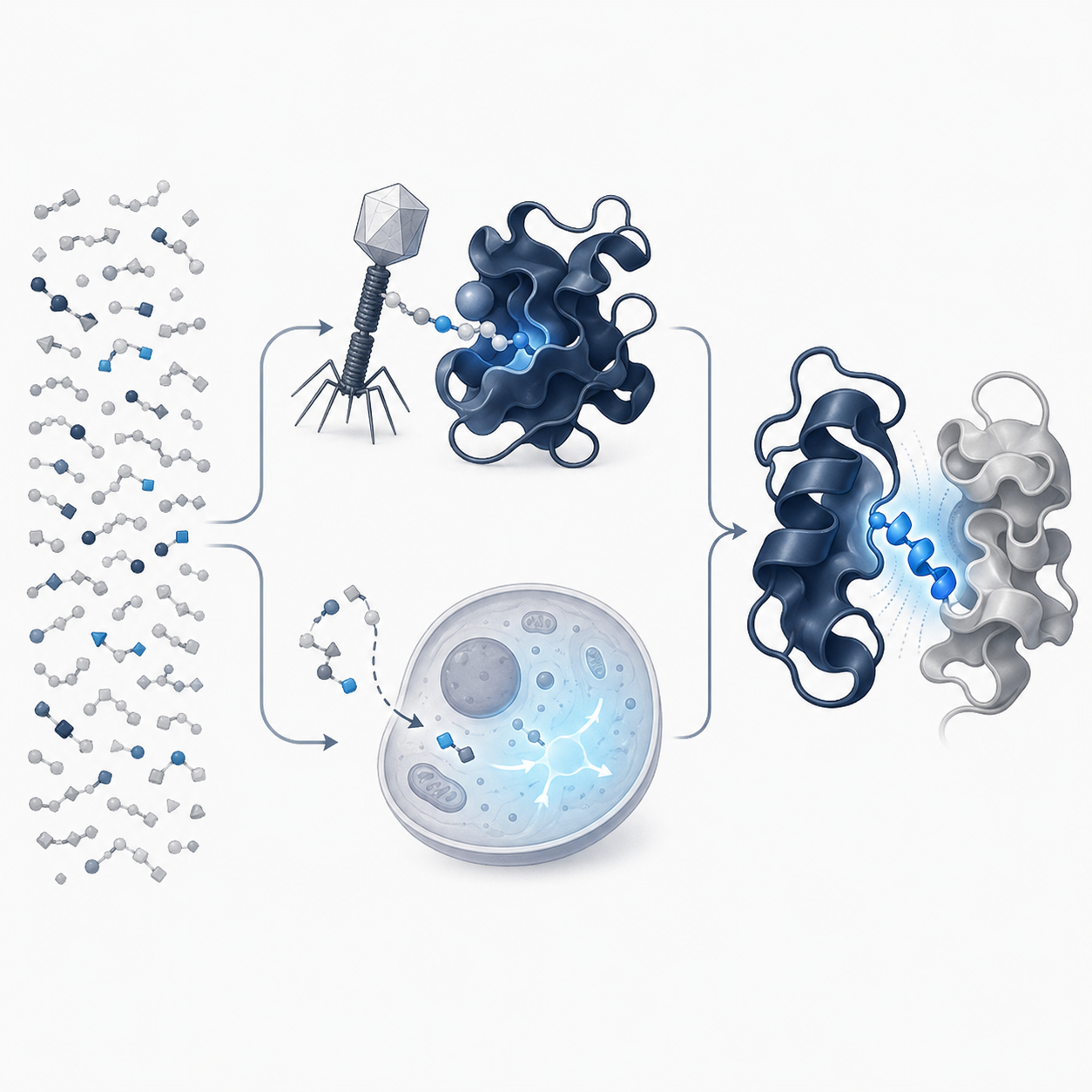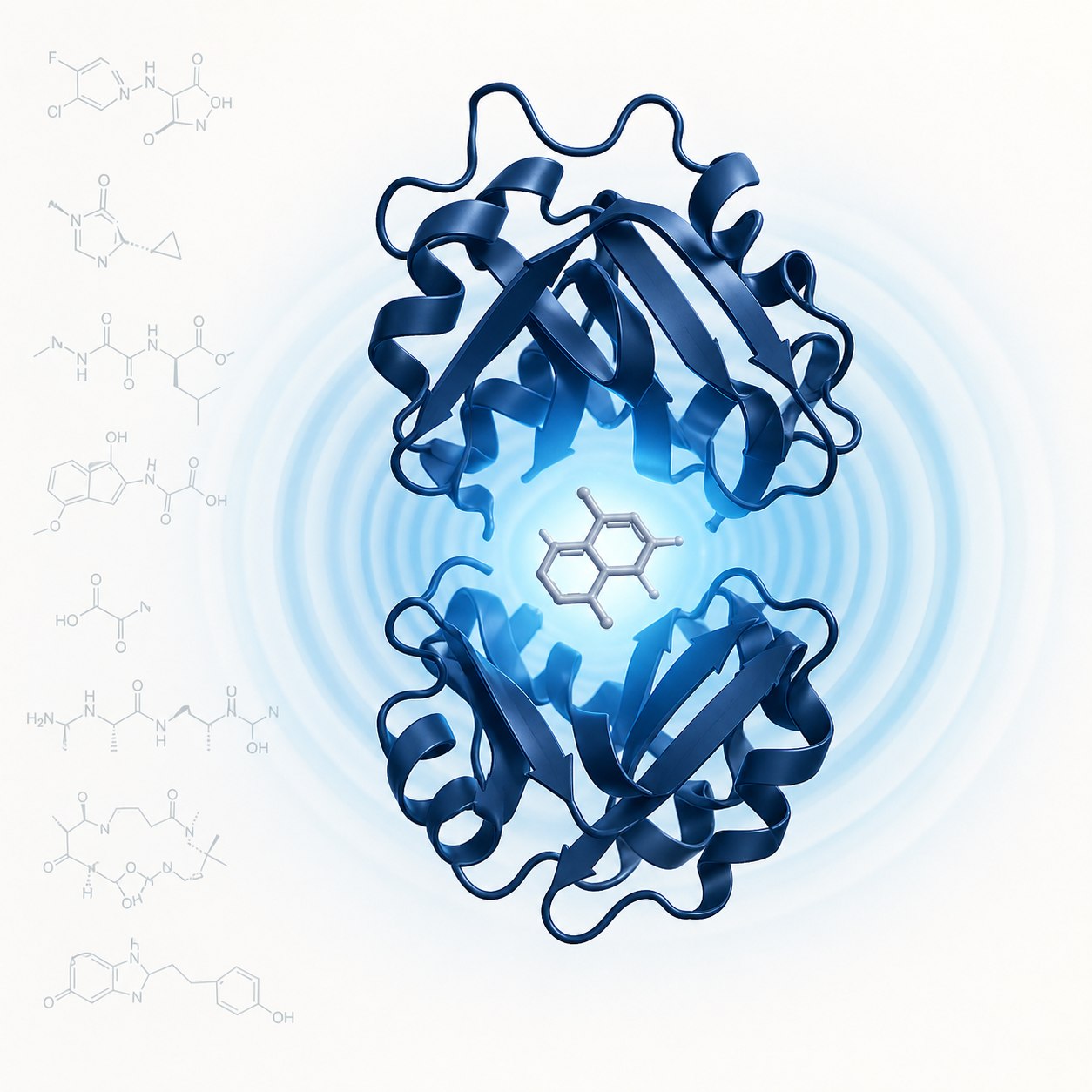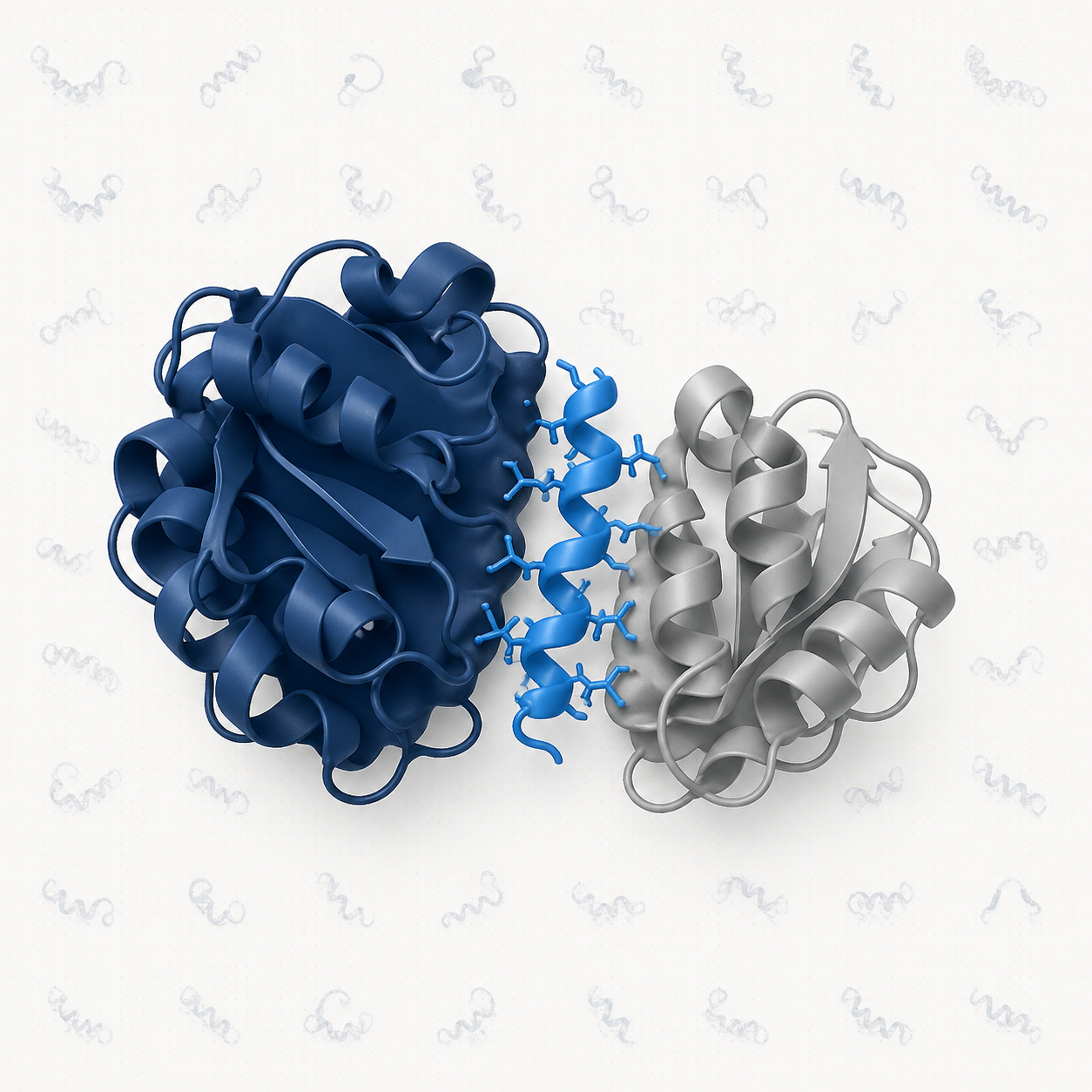
Targeting Protein–Protein Interactions
Protein–protein interactions (PPIs) underlie most cellular processes and represent a vast, largely untapped target space for therapeutic intervention. Yet, traditional drug discovery often misses PPIs because their interfaces are flat and lack well-defined pockets.